Note
Go to the end to download the full example code. or to run this example in your browser via Binder
Custom refit strategy of a grid search with cross-validation#
This examples shows how a classifier is optimized by cross-validation,
which is done using the GridSearchCV object
on a development set that comprises only half of the available labeled data.
The performance of the selected hyper-parameters and trained model is then measured on a dedicated evaluation set that was not used during the model selection step.
More details on tools available for model selection can be found in the sections on Cross-validation: evaluating estimator performance and Tuning the hyper-parameters of an estimator.
# Authors: The scikit-learn developers
# SPDX-License-Identifier: BSD-3-Clause
The dataset#
We will work with the digits dataset. The goal is to classify handwritten
digits images.
We transform the problem into a binary classification for easier
understanding: the goal is to identify whether a digit is 8 or not.
from sklearn import datasets
digits = datasets.load_digits()
In order to train a classifier on images, we need to flatten them into vectors.
Each image of 8 by 8 pixels needs to be transformed to a vector of 64 pixels.
Thus, we will get a final data array of shape (n_images, n_pixels).
n_samples = len(digits.images)
X = digits.images.reshape((n_samples, -1))
y = digits.target == 8
print(
f"The number of images is {X.shape[0]} and each image contains {X.shape[1]} pixels"
)
The number of images is 1797 and each image contains 64 pixels
As presented in the introduction, the data will be split into a training and a testing set of equal size.
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=0)
Define our grid-search strategy#
We will select a classifier by searching the best hyper-parameters on folds of the training set. To do this, we need to define the scores to select the best candidate.
scores = ["precision", "recall"]
We can also define a function to be passed to the refit parameter of the
GridSearchCV instance. It will implement the
custom strategy to select the best candidate from the cv_results_ attribute
of the GridSearchCV. Once the candidate is
selected, it is automatically refitted by the
GridSearchCV instance.
Here, the strategy is to short-list the models which are the best in terms of precision and recall. From the selected models, we finally select the fastest model at predicting. Notice that these custom choices are completely arbitrary.
import pandas as pd
def print_dataframe(filtered_cv_results):
"""Pretty print for filtered dataframe"""
for mean_precision, std_precision, mean_recall, std_recall, params in zip(
filtered_cv_results["mean_test_precision"],
filtered_cv_results["std_test_precision"],
filtered_cv_results["mean_test_recall"],
filtered_cv_results["std_test_recall"],
filtered_cv_results["params"],
):
print(
f"precision: {mean_precision:0.3f} (±{std_precision:0.03f}),"
f" recall: {mean_recall:0.3f} (±{std_recall:0.03f}),"
f" for {params}"
)
print()
def refit_strategy(cv_results):
"""Define the strategy to select the best estimator.
The strategy defined here is to filter-out all results below a precision threshold
of 0.98, rank the remaining by recall and keep all models with one standard
deviation of the best by recall. Once these models are selected, we can select the
fastest model to predict.
Parameters
----------
cv_results : dict of numpy (masked) ndarrays
CV results as returned by the `GridSearchCV`.
Returns
-------
best_index : int
The index of the best estimator as it appears in `cv_results`.
"""
# print the info about the grid-search for the different scores
precision_threshold = 0.98
cv_results_ = pd.DataFrame(cv_results)
print("All grid-search results:")
print_dataframe(cv_results_)
# Filter-out all results below the threshold
high_precision_cv_results = cv_results_[
cv_results_["mean_test_precision"] > precision_threshold
]
print(f"Models with a precision higher than {precision_threshold}:")
print_dataframe(high_precision_cv_results)
high_precision_cv_results = high_precision_cv_results[
[
"mean_score_time",
"mean_test_recall",
"std_test_recall",
"mean_test_precision",
"std_test_precision",
"rank_test_recall",
"rank_test_precision",
"params",
]
]
# Select the most performant models in terms of recall
# (within 1 sigma from the best)
best_recall_std = high_precision_cv_results["mean_test_recall"].std()
best_recall = high_precision_cv_results["mean_test_recall"].max()
best_recall_threshold = best_recall - best_recall_std
high_recall_cv_results = high_precision_cv_results[
high_precision_cv_results["mean_test_recall"] > best_recall_threshold
]
print(
"Out of the previously selected high precision models, we keep all the\n"
"the models within one standard deviation of the highest recall model:"
)
print_dataframe(high_recall_cv_results)
# From the best candidates, select the fastest model to predict
fastest_top_recall_high_precision_index = high_recall_cv_results[
"mean_score_time"
].idxmin()
print(
"\nThe selected final model is the fastest to predict out of the previously\n"
"selected subset of best models based on precision and recall.\n"
"Its scoring time is:\n\n"
f"{high_recall_cv_results.loc[fastest_top_recall_high_precision_index]}"
)
return fastest_top_recall_high_precision_index
Tuning hyper-parameters#
Once we defined our strategy to select the best model, we define the values of the hyper-parameters and create the grid-search instance:
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
tuned_parameters = [
{"kernel": ["rbf"], "gamma": [1e-3, 1e-4], "C": [1, 10, 100, 1000]},
{"kernel": ["linear"], "C": [1, 10, 100, 1000]},
]
grid_search = GridSearchCV(
SVC(), tuned_parameters, scoring=scores, refit=refit_strategy
)
grid_search.fit(X_train, y_train)
All grid-search results:
precision: 1.000 (±0.000), recall: 0.854 (±0.063), for {'C': 1, 'gamma': 0.001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.257 (±0.061), for {'C': 1, 'gamma': 0.0001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.877 (±0.069), for {'C': 10, 'gamma': 0.001, 'kernel': 'rbf'}
precision: 0.968 (±0.039), recall: 0.780 (±0.083), for {'C': 10, 'gamma': 0.0001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.877 (±0.069), for {'C': 100, 'gamma': 0.001, 'kernel': 'rbf'}
precision: 0.905 (±0.058), recall: 0.889 (±0.074), for {'C': 100, 'gamma': 0.0001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.877 (±0.069), for {'C': 1000, 'gamma': 0.001, 'kernel': 'rbf'}
precision: 0.904 (±0.058), recall: 0.890 (±0.073), for {'C': 1000, 'gamma': 0.0001, 'kernel': 'rbf'}
precision: 0.695 (±0.073), recall: 0.743 (±0.065), for {'C': 1, 'kernel': 'linear'}
precision: 0.643 (±0.066), recall: 0.757 (±0.066), for {'C': 10, 'kernel': 'linear'}
precision: 0.611 (±0.028), recall: 0.744 (±0.044), for {'C': 100, 'kernel': 'linear'}
precision: 0.618 (±0.039), recall: 0.744 (±0.044), for {'C': 1000, 'kernel': 'linear'}
Models with a precision higher than 0.98:
precision: 1.000 (±0.000), recall: 0.854 (±0.063), for {'C': 1, 'gamma': 0.001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.257 (±0.061), for {'C': 1, 'gamma': 0.0001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.877 (±0.069), for {'C': 10, 'gamma': 0.001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.877 (±0.069), for {'C': 100, 'gamma': 0.001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.877 (±0.069), for {'C': 1000, 'gamma': 0.001, 'kernel': 'rbf'}
Out of the previously selected high precision models, we keep all the
the models within one standard deviation of the highest recall model:
precision: 1.000 (±0.000), recall: 0.854 (±0.063), for {'C': 1, 'gamma': 0.001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.877 (±0.069), for {'C': 10, 'gamma': 0.001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.877 (±0.069), for {'C': 100, 'gamma': 0.001, 'kernel': 'rbf'}
precision: 1.000 (±0.000), recall: 0.877 (±0.069), for {'C': 1000, 'gamma': 0.001, 'kernel': 'rbf'}
The selected final model is the fastest to predict out of the previously
selected subset of best models based on precision and recall.
Its scoring time is:
mean_score_time 0.004748
mean_test_recall 0.877206
std_test_recall 0.069196
mean_test_precision 1.0
std_test_precision 0.0
rank_test_recall 3
rank_test_precision 1
params {'C': 1000, 'gamma': 0.001, 'kernel': 'rbf'}
Name: 6, dtype: object
The parameters selected by the grid-search with our custom strategy are:
grid_search.best_params_
{'C': 1000, 'gamma': 0.001, 'kernel': 'rbf'}
Finally, we evaluate the fine-tuned model on the left-out evaluation set: the
grid_search object has automatically been refit on the full training
set with the parameters selected by our custom refit strategy.
We can use the classification report to compute standard classification metrics on the left-out set:
from sklearn.metrics import classification_report
y_pred = grid_search.predict(X_test)
print(classification_report(y_test, y_pred))
precision recall f1-score support
False 0.99 1.00 0.99 807
True 1.00 0.87 0.93 92
accuracy 0.99 899
macro avg 0.99 0.93 0.96 899
weighted avg 0.99 0.99 0.99 899
Note
The problem is too easy: the hyperparameter plateau is too flat and the output model is the same for precision and recall with ties in quality.
Total running time of the script: (0 minutes 8.472 seconds)
Related examples

Balance model complexity and cross-validated score

Comparing randomized search and grid search for hyperparameter estimation
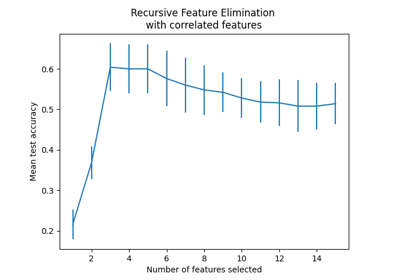
Recursive feature elimination with cross-validation

